Modify multiple elements of a Numpy ndarray with ravel_multi_index
Indexing a multi-dimensional Numpy array
Accessing a multi-dimensional Numpy array by indices is useful for many tasks, such as parsing the contours in a DICOM-RTSTRUCT file.
As a simple example, we are given a 5 x 6 array arr, and a list of the coordinates of 3 points coords:
>>> import numpy as np
>>> arr = np.arange(30).reshape(5,6)
>>> coords = np.array([[0, 1], [3, 4], [3, 2]])
>>> arr
array([[ 0, 1, 2, 3, 4, 5],
[ 6, 7, 8, 9, 10, 11],
[12, 13, 14, 15, 16, 17],
[18, 19, 20, 21, 22, 23],
[24, 25, 26, 27, 28, 29]])
>>> coords
array([[0, 1],
[3, 4],
[3, 2]])
To obtain the values of the targeted pixels as a list [arr[0,1], arr[3,4], arr[3,2]], that is [1, 22, 20], it is tempting to write “arr[coords]“. But this is incorrect.
>>> arr[coords]
array([[[ 0, 1, 2, 3, 4, 5],
[ 6, 7, 8, 9, 10, 11]],
[[18, 19, 20, 21, 22, 23],
[24, 25, 26, 27, 28, 29]],
[[18, 19, 20, 21, 22, 23],
[12, 13, 14, 15, 16, 17]]])
Using a list of indices like this is correct in MATLAB, but not in Numpy. Why?
Fancy indexing
Passing an array of indices to access multiple elements at once is referred to as Fancy Indexing in Numpy.
The most important rule to remember is that:
The (broadcasted) shape of the index array determines that of the output array.
>>> coords.shape
(3, 2)
>>> arr[coords].shape
(3, 2, 6)
Fancy indexing is basically a way to avoid writing Python loops, by having numpy do it internally for you with its more efficient, lower-level loops.
In this case, arr[coords] is actually sampling the rows rather than the elements. Conceptually, we get:
array([[arr[0], arr[1]],
[arr[3], arr[4]],
[arr[3], arr[2]]])
or
array([[0th row, 1st row],
[3rd row, 4th row],
[3rd row, 2nd row]])
The correct way is to first obtain both the column and row indices as two lists (1D arrays), and then pass them to arr for fancy indexing.
>>> rows = coords[:,0]
>>> cols = coords[:,1]
>>> arr[rows, cols]
array([ 1, 22, 20])
numpy.unravel_index
Another approach would be to access the multi-dimensional array as a 1D array.
In C++, accessing a multi-dimensional array can be done as though it were a 1D array: “arr[col][row]” is equivalent to “arr[(row-1)*n_cols + col]“. This is because that multi-dimensional array is allocated on a contiguous span of memory, which is 1D. “(row-1)*n_cols + col” is basically how deep you’re into this span of memory.
The internal data representation of a Numpy array is conceptually the same. You can use ravel() to obtain a 1D view of the array without copying, and use “ravel_multi_index()” to translate the indices into 1D.
ravel_multi_index() and numpy.unravel_index() are inverse operations. Their MATLAB equivalents are sub2ind and ind2sub.
>>> arr.ravel()[np.ravel_multi_index(coords.T, arr.shape)]
array([ 1, 22, 20])
Modify multiple elements at once
It is worth noting that, using either fancy indexing method, not only you can read out the elements, but also modify them at once.
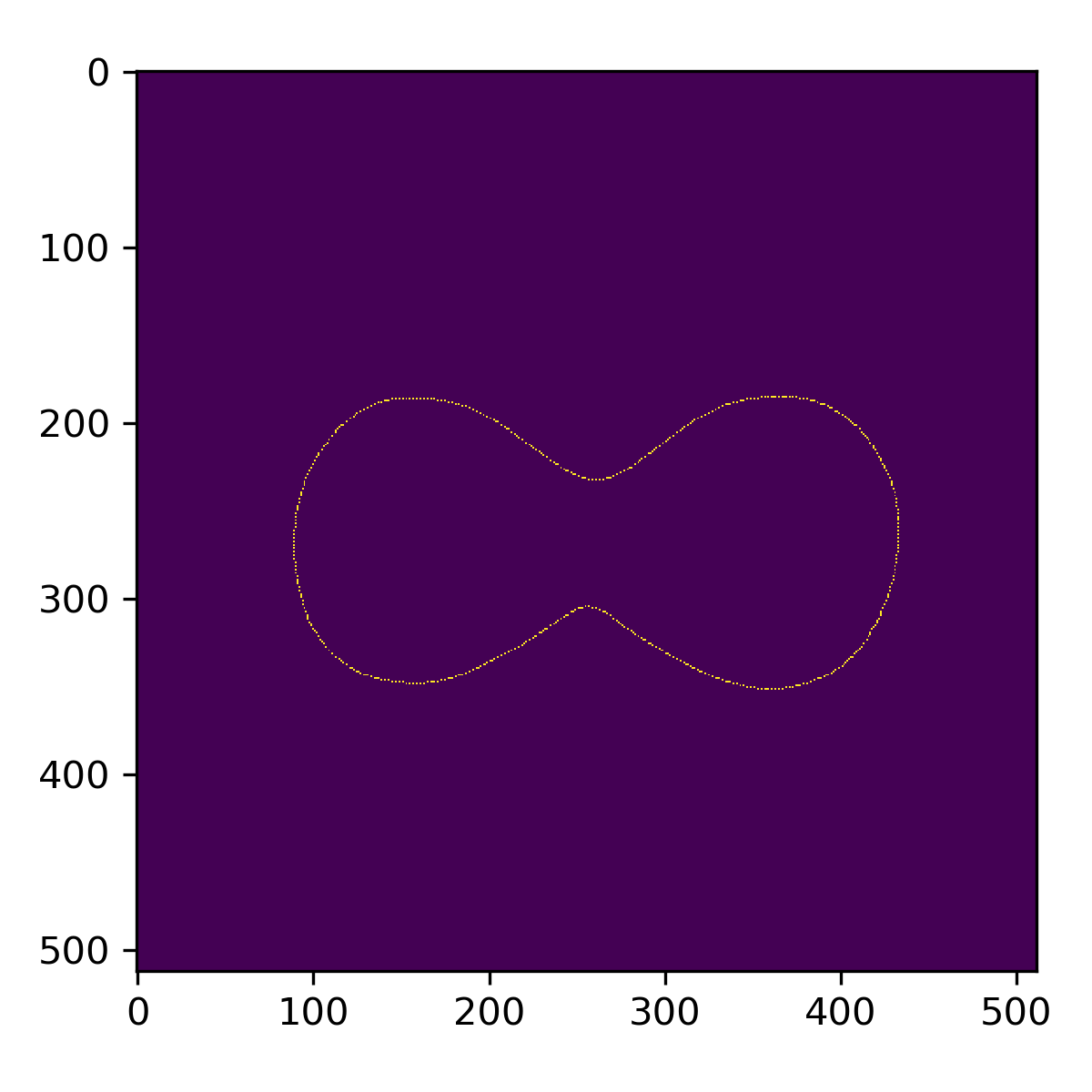
This is particularly useful if you need to parse the discrete contour points into a 2D map like below.
>>> img = np.zeros((5,6)).astype(bool)
>>> img
array([[False, False, False, False, False, False],
[False, False, False, False, False, False],
[False, False, False, False, False, False],
[False, False, False, False, False, False],
[False, False, False, False, False, False]])
>>> img[rows, cols] = True
>>> img
array([[False, True, False, False, False, False],
[False, False, False, False, False, False],
[False, False, False, False, False, False],
[False, False, True, False, True, False],
[False, False, False, False, False, False]])
or
>>> img.ravel()[np.ravel_multi_index(coords.T, img.shape)] = True
>>> img
array([[False, True, False, False, False, False],
[False, False, False, False, False, False],
[False, False, False, False, False, False],
[False, False, True, False, True, False],
[False, False, False, False, False, False]])